Dana-Farber/Harvard Cancer Center SPORE in Gastrointestinal Cancer
Dana-Farber Cancer Institute
Principal Investigator(s):

William R. Sellers, MD

Nabeel Bardeesy, PhD
- Principal Investigator(s) Contact Information
- Overview
- Project 1: Integrating Targeted and Immune Therapies for BRAF-Mutant Colorectal Cancers
- Project 2: Exploiting FGF Receptor Signaling Dependence in Liver Cancer Therapy
- Project 3: Improving Therapy for DNA-Damage Deficient Pancreatic Adenocarcinoma
- Project 4: Characterizing and Overcoming Resistance to ERBB2 Directed Therapy in Metastatic Gastric and Esophageal Adenocarcinoma
- Administrative Core
- Pathology and Biospecimen Core
- Bioinformatics and Biostatistics Core
- Developmental Research Program
- Career Enhancement Program
Principal Investigator(s) Contact Information
William R. Sellers, MD
Director of the Cancer Program
Broad Institute of MIT & Harvard
Faculty Member at Harvard Medical School
Faculty Member at Dana-Farber Cancer Institute
415 Main St., Office 4009
Cambridge, MA 02142
617-714-7110
Nabeel Bardeesy, PhD
Massachusetts General Hospital
Simches Research Building
185 Cambridge Street, CPZN-420
Boston, MA 02114
(617) 643-2579
Overview
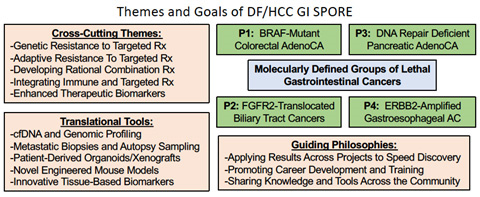
This SPORE brings together the respective strengths and investigators from multiple hospitals and research centers within the Harvard Medical Community with the goal of linking science and medicine to advance our understanding of and treatment for gastrointestinal cancers. While gastrointestinal malignancies are a diverse set of diseases, there are key biologic and clinical problems that transcend these diseases. Furthermore, beyond addressing these specific biologic problems, our SPORE aims to establish core resources that enhance our ability to perform translational research into gastrointestinal cancers and to support new developmental research projects and career development awards that spark new ideas and help establish the careers of new generations of researchers.
To address the need to advance therapy for these diseases by building upon fundamental disease biology, we have established a set of projects across distinct GI cancers where therapy remains largely reliant upon empiric chemotherapy. To make progress in this field, we have designed our projects to build on the discovery of molecularly distinct subsets of GI cancer subtypes for a which a targeted therapeutic approach is showing promise. In each case, however, the clinical impact of these new therapeutic avenues has been limited by uncertainty about patient selection and limited biomarker development, by insufficient understanding of tumor circuitry, and by intrinsic and acquired modes of drug resistance, including immune evasion. The projects will thus create paradigms for subsequent rational, mechanism-based attack across GI cancers. Furthermore, by studying rational targeted therapies in the GI system in our collaborative community, the impact of these projects is enhanced as results from each project can be readily evaluated across projects.
Project 1: Integrating Targeted and Immune Therapies for BRAF-Mutant Colorectal Cancers
Project Co-Leaders:
Ryan Corcoran, MD, PhD (clinical co-leader)
Arlene Sharpe, MD, PhD (basic co-leader)
Here we will explore potential cooperativity between targeted MAPK inhibition (MAPKi) and immune checkpoint blockade (ICB) to convert less immune responsive tumors to more immunogenic tumors. BRAFm CRC represents a prime population for exploring potential cooperativity, as 20-30% of metastatic BRAFm CRCs harbor MSI, which confers responsiveness to ICB. Moreover, we have observed durable responses of >5 years in MSI BRAFm CRC patients receiving MAPKi alone. We propose a comprehensive effort using innovative immune competent BRAFm CRC mouse models, cutting-edge molecular and immune analyses of paired pre- and on-treatment tumor biopsies, and novel clinical trials to explore combined MAPKi and ICB as a strategy to achieve durable benefit in BRAFm CRC patients.
Project 2: Exploiting FGF Receptor Signaling Dependence in Liver Cancer Therapy
Project Co-Leaders:
Nabeel El-Bardeesy, PhD (basic co-leader)
Cyril Benes, PhD (basic co-leader)
Andrew Zhu, MD, PhD (clinical co-leader)
Intrahepatic cholangiocarcinoma (ICC) ranks among the deadliest cancers, with a 5-year survival rate of only 5-15%. The discovery that receptor tyrosine kinase Fibroblast Growth Factor Receptor 2 (FGFR2) is genetically activated in >20% of ICCs, most commonly via FGFR2 gene fusions, is leading to a new therapeutic paradigm for a subset of patients with this disease. Indeed, early clinical trials with FGFR inhibitors (FGFRi) show significant anti-tumor efficacy. We propose a multi-pronged strategy for impactful translational research that leverages serial tumor and liquid biopsies from three FGFRi trials, a series of patient-derived xenografts (PDX) and organoids, a genetically engineered mouse model, and novel FGFR2 inhibitors. We will systematically address the key barriers to more effective targeted therapy against FGFR2 in ICC: 1) the mechanism of FGFRi resistance, either due to acquisition of secondary FGFR2 mutations or activation of ‘bypass’ signaling pathways, 2) identification of more effective FGFR2 inhibitors, including those that can target the kinase after it acquires resistance mutations, and 3) defining novel combination therapies to target bypass pathways to counter resistance and prolong initial response.
Project 3: Improving Therapy for DNA-Damage Deficient Pancreatic Adenocarcinoma
Project Co-Leaders:
Alan D’Andrea, MD (basic co-leader)
Geoff Shapiro, MD, PhD (basic co-leader)
Brian Wolpin, MD (clinical co-leader)
Pancreatic ductal adenocarcinoma (PDAC) is the third leading cause of cancer-related death in the United States. Up to 20% of PDAC patients harbor mutations in genes responsible for double-strand DNA damage repair (DDR), and these defects may unveil vulnerabilities to treatment with a novel class of drugs called PARP inhibitors. A subset of PDAC patients with mutations in BRCA1 and BRCA2, as well as other DDR genes, may have durable tumor responses to poly(ADP-ribose) polymerase (PARP) inhibitors; however, the optimal biomarkers have not been identified to predict which patients will benefit from these therapies. Furthermore, combination treatment programs to move beyond single-agent PARP inhibition are not yet defined. This proposal brings together a team of distinguished laboratory, translational and clinical investigators to: (1) define optimal genomic and functional strategies for identifying PDAC patients with DDR deficiency; (2) conduct treatment trials to identify the patients with greatest benefit from PARP inhibition and to identify mechanisms of de novo and acquired resistance; and (3) to define novel combination treatment strategies for future clinical trials.
Project 4: Characterizing and Overcoming Resistance to ERBB2 Directed Therapy in Metastatic Gastric and Esophageal Adenocarcinoma
Project Co-Leaders:
Adam Bass, MD (basic co-leader)
Peter Enzinger, MD (clinical co-leader)
ERBB2 is amplified in ~20% of Gastric and esophageal adenocarcinomas (GEAs) and metastatic ERBB2+ GEAs are treated with a combination of chemotherapy and the antibody Trastuzumab. However, Trastuzumab is only modestly effective in GEA, and all other targeted agents in ERBB2+ breast cancer have failed in GEA clinical trials. We propose to directly address the two primary factors that we hypothesize to mediate failure of ERBB2 therapy in GEA: adaptive resistance and genetic complexity. To perform these studies, our team of investigators with complementary skill sets will both perform detailed assessment of optimal approaches to stably inhibit ERBB2 using an array of patient-derived model systems. Furthermore, we will perform a prospective clinical collection spanning multiple large academic medical centers in which we evaluate the genetic evolution of ERBB2+ GEAs during therapy, define genetic alterations that accompany resistance and then functionally validate mechanisms of resistance and optimal combination therapy. We will also explore the role of cell-free (cf)DNA genomic profiling to guide therapy in the face of genomic evolution of the disease during therapy. The overall goal will be to validate candidate resistance mechanisms and seek to define optimal combination therapies that can overcome them.
Administrative Core
Core Directors:
William R. Sellers, MD (Co-Director)
Nabeel Bardeesy, PhD (Co-Director)
The purpose of the Administration, Evaluation and Planning Core is to coordinate the varied components of the DF/HCC GI SPORE necessary to provide oversight and leadership of the scientific, administrative and fiscal aspects of the SPORE.
Pathology and Biospecimen Core
Core Directors:
Jason Hornick, MD (Co-Director)
Vikram Deshpande (Co-Director)
The Biospecimen and Pathology Core will work in close collaboration with investigators to achieve three crucial purposes: i) to provide tissue specimens and pathology expertise to evaluate human and mouse models of GI neoplasms; ii) to provide molecular pathologic expertise in state-of-the-art molecular evaluation of tissue specimens; and iii) to evaluate and implement new tissue and cellular imaging and analysis technologies that will benefit these research goals in the future.
Bioinformatics and Biostatistics Core
Core Directors:
Dianne Finkelstein, PhD (Co-Director)
Gad Getz, PhD (Co-Director)
The Biostatistics and Bioinformatics Core includes leading quantitative research scientists who will work with SPORE personnel to assure that the research carried out meets the prevailing and emerging research standards and adapts as new technologies develop and are introduced into the scientific research program. This support will be provided by establishing, maintaining, and supporting collaborative relationships that will ensure that Core members are integrated members of the research teams and provide appropriate statistical and computational support for all GI cancer SPORE investigators. This will include consultation and collaboration on all aspects of study design, database development and quality control, and analysis and interpretation of data.
Developmental Research Program
Program Directors:
Ramesh Shivdasani, MD, PhD
Debroah Schrag, MD, MPH
Career Enhancement Program
Program Directors:
Robert Mayer, MD
Wendy Garrett, MD, PhD







